Due: Friday, May 10 at 11:59pm
Submission: On Courseworks
Skeleton Code: hw7.zip
Please read the instructions carefully. An incorrectly formatted submission may not be counted.
In this homework, we will start from some skeleton code. You are given the mlmodel module. We will implement part 4 of this homework on top of the skeleton code, just as in homework 6. To submit, place these modules inside a folder called uni-hw7 (for for me this would be tkp2108-hw7). Then compress this folder into a .zip file. For most operating systems, you should be able to right click and have a “compress” option in the context menu. This should create a file called tkp2108-hw7.zip (with your uni). Submit this on courseworks.
Answer the following questions in a file part1.txt:
What is Vectorization? Provide a definition and provide an example of vectorization from class or the homework.
What is recursion? What is required in order to write a recursive algorithm? Give an example of a recursive function from class or the homework.
Provide a brief (1 sentence) explanation of each of the following terms. Then provide an example of the term from the homeworks.
You will find a CSV file with generated data here. Using this file, implement the following python functions with Pandas:
def readData(filename):
"""return a dataframe with index 'Row ID'"""
def filterBy(df, quantity=None, discount=None, profit=None):
"""given the input dataframe, return just the rows where Quantity column is greater than the input variable quantity, and similarly for discount and profit.
If the value passed in is None, do not filter with it.
"""
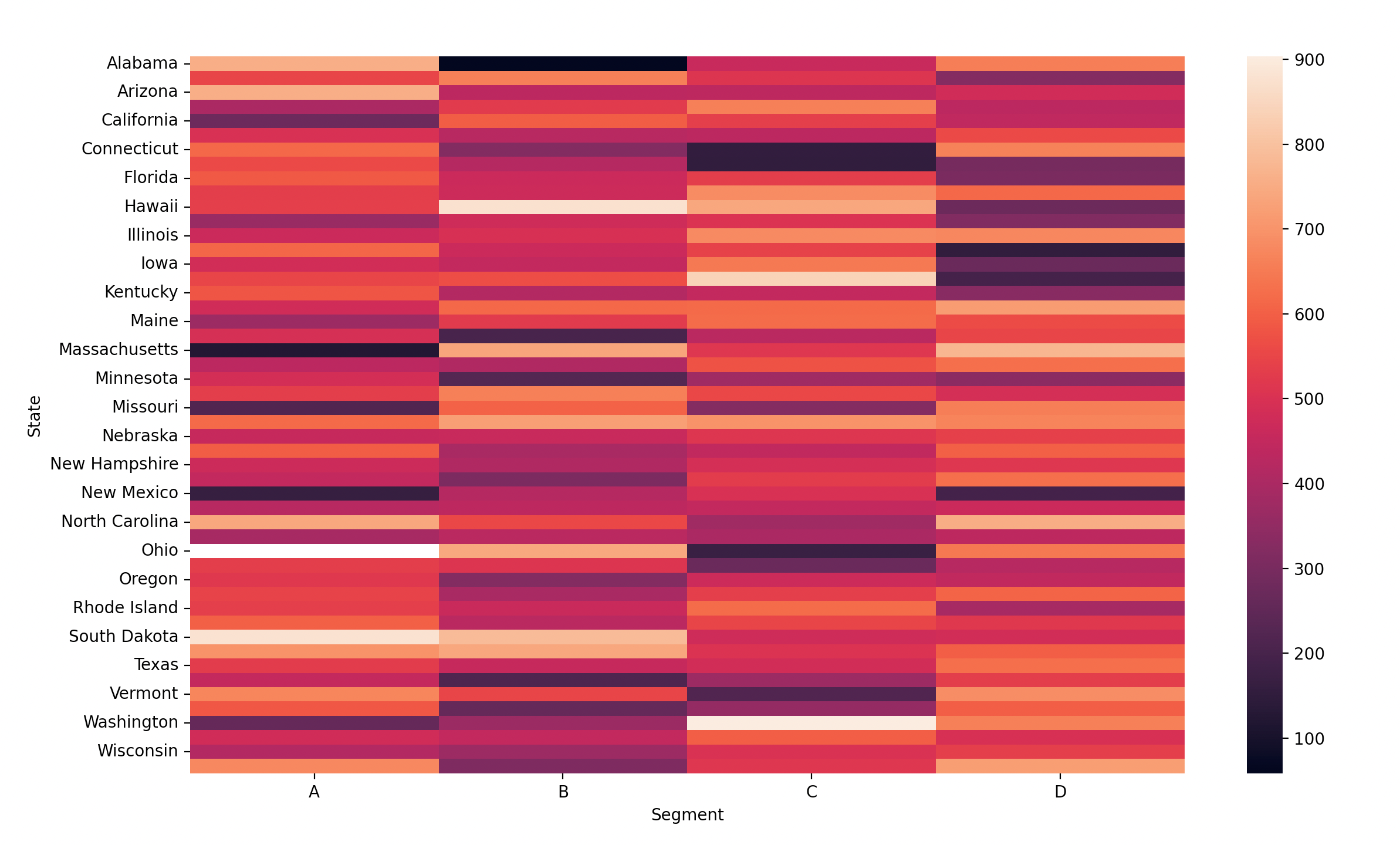
def stateSegmentHeatmap(df)
"""Draw a heatmap with seaborn where:
- states as row pivots
- segments as column pivots
- color by profit
See the example below"""
Implement a new class RandomList that inherits from Python’s standard list, with the following features:
RandomList from that data shuffledRandomList from that number of random integers between 0 and 100__str__ and __repr__ methods to exhibit the behavior below__add__ operator to exhibit the behavior below
list, return a list with the two sets of dataRandomList, return a new RandomList with the concatenated data shuffledshuffle which randomly shuffle’s the RandomList’s dataHere is an example IPython session with with a sample run:
In [1]: rl1 = RandomList(10) # creates a RandomList with 10 random integers between 0 and 100
In [2]: rl2 = RandomList([1, 2, 3]) # creates a RandomList from this list, shuffling the contents
In [3]: rl3 = RandomList("test") # throws error
---------------------------------------------------------------------------
NotImplementedError Traceback (most recent call last)
----> 1 rl3 = RandomList("test") # throws error
final.py in __init__(self, data)
---> 18 raise NotImplementedError
In [4]: print(rl1)
random([28, 31, 22, 63, 45, 8, 28, 9, 31, 96])
In [5]: print(rl2)
random([2, 1, 3])
In [6]: lst = [rl1, rl2]
In [7]: print(lst)
[<RandomList [28, 31, 22, 63, 45, 8, 28, 9, 31, 96]>, <RandomList [2, 1, 3]>]
In [8]: print(rl1)
random([28, 31, 22, 63, 45, 8, 28, 9, 31, 96])
In [9]: rl1.shuffle()
In [10]: print(rl1)
random([63, 9, 96, 28, 8, 22, 31, 45, 28, 31])
In [11]: print(rl1 + [1, 2, 3]) # make new `list` from rl1's data and the other list
[63, 9, 96, 28, 8, 22, 31, 45, 28, 31, 1, 2, 3]
In [12]: print(rl1 + rl2) # make a new RandomList frp, rl1'1 data and the other list, shuffling
random([22, 96, 3, 28, 31, 1, 45, 8, 31, 9, 2, 28, 63])
In [13]: print(rl1 + "abc") # throws error
---------------------------------------------------------------------------
NotImplementedError Traceback (most recent call last)
<ipython-input-14-4481c54f83fc> in <module>
----> 1 print(rl1 + "abc") # throws error
final.py in __add__(self, other)
---> 71 raise NotImplementedError
For this part, we will build on the code we started in hw6 and implement 3 predictive models for solving our supervised classification problem.
As a high level overview, we will do a few things:
This sounds complicated, but in code its pretty straightforward:
# split dataset into training and testing data, with separate labels
# make sure data is randomly shuffled beforehand!!
train, train_labels, test, test_labels = splitDataset(df, test_percentage=20)
# Print some info about test/train split
print("Test dataset has {} entries".format(len(test)))
print("Train dataset has {} entries".format(len(train)))
# run our knn classifier
nn_accuracy = NNClassifier(train, test, train_labels, test_labels, k)
# run the knn classifier from scikit-learn
knn_accuracy = SKLearnKnnClassifier(train, test, train_labels, test_labels, k)
# run the svm classifier from scikit-learn
svm_accuracy = SKLearnSVMClassifier(train, test, train_labels, test_labels)
# print the accuracies
print("Accuracies:\n\tKNN (1006):{:.1%}\n\tKNN (sklearn):{:.1%}\n\tSVM (sklearn):{:.1%}".format(nn_accuracy, knn_accuracy, svm_accuracy))
This should lead to some output like:
Accuracies:
KNN (1006):91.7%
KNN (sklearn):91.7%
SVM (sklearn):89.3%
More concretely, we will be implementing 3 main functions:
Recall from class the appeal of scikit-learn: Polymorphism! That means that all scikit-learn classifiers have the same shape:
model.fit(training_data, training_labels)
predicted_labels = model.predict(testing_data)
So both these functions should be super straightforward to implement
def SKLearnKnnClassifier(training, testing, training_labels, testing_labels, k):
# instantiate model
knn = KNeighborsClassifier(n_neighbors=k)
# fit model to training data
# predict test labels
# return % where prediction matched actual
def SKLearnSVMClassifier(training, testing, training_labels, testing_labels):
# instantiate model
svm = SVC(kernel="rbf")
# fit model to training data
# predict test labels
# return % where prediction matched actual
This function is our handwritten K Nearest Neighbors algorithm. It sounds a littel daunting, but in reality its pretty straightforward. My skeleton code breaks it apart into two separate steps:
knn function to get the predicted labelLet’s start with the first function:
def NNClassifier(training, testing, training_labels, testing_labels, k):
# preallocate labels
predicted_labels = np.zeros(len(testing_labels)).astype(str)
# run knn on each point and assign its label
# fill in the labels with knn
# return % where prediction matched actual
return 0.0
For each row in testing, we call knn to get back a label ("B" or "M") and we put that in a preallocated vector predicted_labels. Then at the end, we compare predicted_labels to testing_labels to generate our accuracy. When we’re testing NNClassifier, we can have knn just hardcoded to return “B” or “M” until we implement it.
def knn(data, data_labels, vector, k):
return "B"
For the knn function, we have to do a little work. Given known datapoints data with labels data_labels, and unknown data point vector and an input parameter k:
vector to each known data point in datakdata_labels corresponding to those kdef knn(data, data_labels, vector, k):
# preallocate distance array
distances = np.zeros(len(data_labels))
# calculate distances
# set labels
# take vote amongs top labels
Here is one possible solution (but you’re welcome to come up with your own!)
To calculate euclidean distances, we can use exactly the same strategy as from our nbody example (just in 30 dimensions). For each data point in data, calculate the distance between it and vector and put the result into the preallocated array distances.
To find the closest k, we just want to sort from minimum to maximum. But be careful! If we try to sort our distances array, we’ll lose the order, and we need the order because each position in distances corresponds to a point in data and a label in data_labels. So if we sort distances, the element at position 5 might move to position 3, but the label in data_labels will still be at position 5! So we need to be careful not to reorder our data.
Instead, we can take advantage of some clever numpy slicing. Recall the following code:
x = np.array([5, 2, 1, 4, 0])
# we can slice with an array of positions to select
x[ [0, 1, 2] ] # slice the 0th position, 1st position, and 2nd position element
# result: array([5, 2, 1])
So if we could calculate the order of the elements without rearranging them:
x = np.array([3, 5, 3, 2, 2, 1, 4, 4, 0])
order = some_function(x)
# order == [8, 5, 3, 4, 0, 2, 6, 7, 1]
# because the 8th element of x (0) would be 1st in order if sorted
# the 5th element of x (1) would be 2nd in order if sorted
# the 3rd element of x (2) would be 3rd in order if sorted
# the 4th element of x (2) would be 4th in order if sorted
Then slicing the data would put it in order:
x = np.array([3, 5, 3, 2, 2, 1, 4, 4, 0])
order = some_function(x)
x[order]
# result: [0, 1, 2, 2, 3, 3, 4, 4, 5]
But with our code, if we can calculate the order of the distances, then we can slice the labels with the same order:
distances = np.array([0, 3, 2, 0, 1])
labels = np.array(["B", "M", "M", "B", "B"])
order = some_function(distances) # [0, 3, 4, 2, 1]
labels_in_order = labels[order] # ['B', 'B', 'B', 'M', 'M']
And to look at the k nearest labels, we just look at the first k labels of labels_in_order!
k = 3
first_k_labels = labels_in_order[:k] # ["B", "B", "B"]
predicted_label = mode(first_k_labels) # mode(["B", "B", "B"]) which is "B"